Kimitoshi KIMURA
Program-Specific Junior Associate Professor, Department of Therapeutics for Multiple System Atrophy, Kyoto University Graduate School of Medicine, Japan
Kyoto University FOREST Principal Investigator
Lab page
Fields of Expertise
Microglia; neurodegeneration of the central nervous system; multiple sclerosis; Alzheimer’s disease
Research Overview
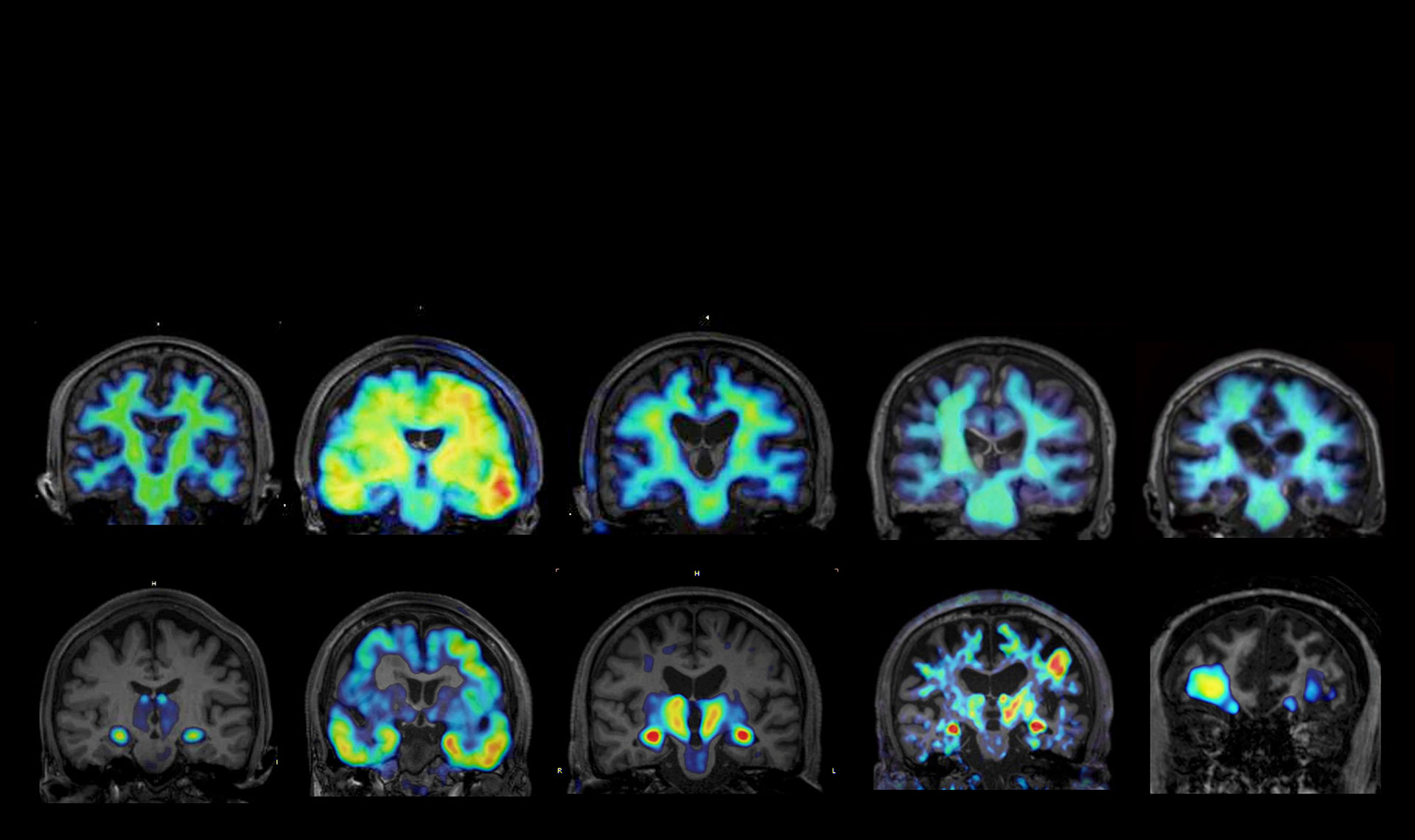
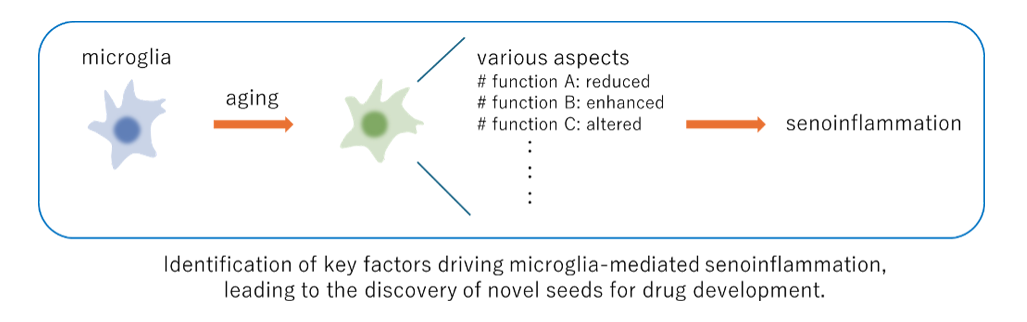
I lead a subproject entitled “Comprehensive screening and functional characterization of key mediators of microglial senoinflammation,” in Moonshot Goal 7, “Early detection and modulation of the dementia pathogenesis based on the concept evolving from glial pathology to senoinflammation.” Microglia play a critical role in central nervous system aging and are deeply involved in various neurodegenerative disorders, including Alzheimer’s disease. Regulation of age-associated inflammatory phenotypes of microglia—contributing to senoinflammation—holds promise for therapeutic intervention not only in aging itself but also in a wide range of refractory neurological diseases. However, the precise molecular mechanisms governing microglial regulation remain incompletely understood. In this project, we aim to comprehensively search for key factors involved in microglial senoinflammatory phenotypes. In parallel, we will construct a gene expression database that captures the multifaceted functions of microglia, with a particular focus on senoinflammatory aspects, enabling further refinement of candidate regulators of microglial senoinflammation. Ultimately, our goal is to identify molecular targets that can serve as seeds for drug discovery to modulate central nervous system aging.
Main Research Topics
1. Comprehensive screening of gene sets involved in microglial senoinflammation
We will perform a comprehensive analysis of microglial phenotypes associated with aging-related inflammation in the central nervous system. By leveraging existing microglial gene expression datasets, we will identify gene sets that show characteristic changes during aging. Among these aging-associated genes, we will use bioinformatics approaches to infer hub genes that are likely to play central functional roles. Candidate genes implicated in microglial aging will then be subjected to screening using primary mouse microglia and microglial cell lines to identify key regulators of microglial senoinflammation.
2. Identification of promising drug targets through experimental validation of candidate key mediators of microglial senoinflammation
Aging-related genes identified through the above screening will be individually validated through experimental analyses. Functional studies, including investigation of intracellular mechanisms, will be conducted using mouse primary microglia and microglial cell lines. In addition, human iPS cell–derived microglia will be employed, using changes in aging-related gene expression as readouts for experimental validation. Through these approaches, we aim to identify target molecules whose modulation can reverse or alleviate microglial aging, thereby serving as candidate drug targets.
3. Construction of a comprehensive gene expression guide focusing on senoinflammatory aspects of microglia
Using existing microglial gene expression datasets, we will define gene expression profiles that are altered in aging and inflammatory states of microglia. Beyond aging- and inflammation-associated profiles, we will also identify multiple additional profiles that underlie the functional heterogeneity of microglia. This integrative analysis will clarify gene expression signatures and intracellular signaling pathways specific to senoinflammation. Based on these findings, we will construct a comprehensive guide to microglial gene expression profiles, which will be shared and utilized in related research projects both within and beyond the Moonshot program.
Publication
Kimura K, Subramanian A, Yin Z, Khalilnezhad A, Wu Y, He D, Dixon KO, Chitta UK, Ding X, Adhikari N, Guzchenko I, Zhang X, Tang R, Pertel T, Myers SA, Aastha A, Nomura M, Eskandari-Sedighi G, Singh V, Liu L, Lambden C, Kleemann KL, Gupta N, Barry JL, Durao A, Cheng Y, Silveira S, Zhang H, Suhail A, Delorey T, Rozenblatt-Rosen O, Freeman GJ, Selkoe DJ, Weiner HL, Blurton-Jones M, Cruchaga C, Regev A, Suvà ML, Butovsky O, Kuchroo VK. Immune checkpoint TIM-3 regulates microglia and Alzheimer’s disease. Nature. 2025, 641(8063):718-731.
Nishigori R, Hamatani M, Yoshitomi H, Kimura K, Takata M, Ashida S, Fujii C, Ochi H, Takahashi R, Kondo T, Ueno H. CD21lo B cell subsets are recruited to the central nervous system in acute neuromyelitis optica. Brain. 2025, awaf086.
Kimura K. Role of resident memory T cells in neuroinflammatory and neurodegenerative diseases in the central nervous system. Neural Regen Res. 2025, 20(11):3227-3228.
Hou Y, Sun L, LaFleur MW, Huang L, Lambden C, Thakore PI, Geiger-Schuller K, Kimura K, Yan L, Zang Y, Tang R, Shi J, Barilla R, Deng L, Subramanian A, Wallrapp A, Choi HS, Kye YC, Ashenberg O, Schiebinger G, Doench JG, Chiu IM, Regev A, Sharpe AH, Kuchroo VK. Neuropeptide signalling orchestrates T cell differentiation. Nature. 2024, 635(8038):444-452.
Kimura K, Nishigori R, Hamatani M, Sawamura M, Ashida S, Fujii C, Takata M, Lin Y, Sato W, Okamoto T, Kuzuya A, Takahashi R, Yamamura T, Kondo T. Resident Memory-like CD8+ T Cells Are Involved in Chronic Inflammatory and Neurodegenerative Diseases in the CNS. Neurol Neuroimmunol Neuroinflamm. 2024, 11(1), e200172.
Kimura K, Lin Y, Yamaguchi H, Sato W, Takewaki D, Minote M, Doi Y, Okamoto T, Takahashi R, Kondo T, Yamamura T. Th1 – CD11c+ B Cell Axis Associated with Response to Plasmapheresis in Multiple Sclerosis. Annals of Neurology, 2021, 90(4), 595-611.
Kimura K. Regulatory T cells in multiple sclerosis. Clinical and Experimental Neuroimmunology, 2020, 11(3), 148-155.
Kimura K, Okada Y, Fujii C, Komatsu K, Takahashi R, Matsumoto S, Kondo T. Clinical characteristics of autoimmune disorders in the central nervous system associated with myasthenia gravis. Journal of Neurology, 2019, 266(11), 2743-2751.
Kimura K, Hohjoh H, Fukuoka M, Sato W, Oki S, Tomi C, Yamaguchi H, Kondo T, Takahashi R, Yamamura T. Circulating exosomes suppress the induction of regulatory T cells via let-7i in multiple sclerosis. Nature Communications, 2018, 9(1), 17.
Kimura K, Hohjoh H, Yamamura T. The Role for Exosomal microRNAs in Disruption of Regulatory T Cell Homeostasis in Multiple Sclerosis. Journal of Experimental Neuroscience, 2018, 12, 1179069518764892.
Kimura K, Yamamura T. Circulating exosomes cause loss of immunological balance in multiple sclerosis. Clinical and Experimental Neuroimmunology, 2018, 9(4), 12474.
Kimura K, Nakamura M, Sato W, Okamoto T, Araki M, Lin Y, Murata M, Takahashi R, Yamamura T. Disrupted balance of T cells under natalizumab treatment in multiple sclerosis. Neurol Neuroimmunol Neuroinflamm. 2016, 3(2), e210.