研究室リンク
Field of Expertise
Systems Biology, Bioinformatics, Deep Learning
Research Overview
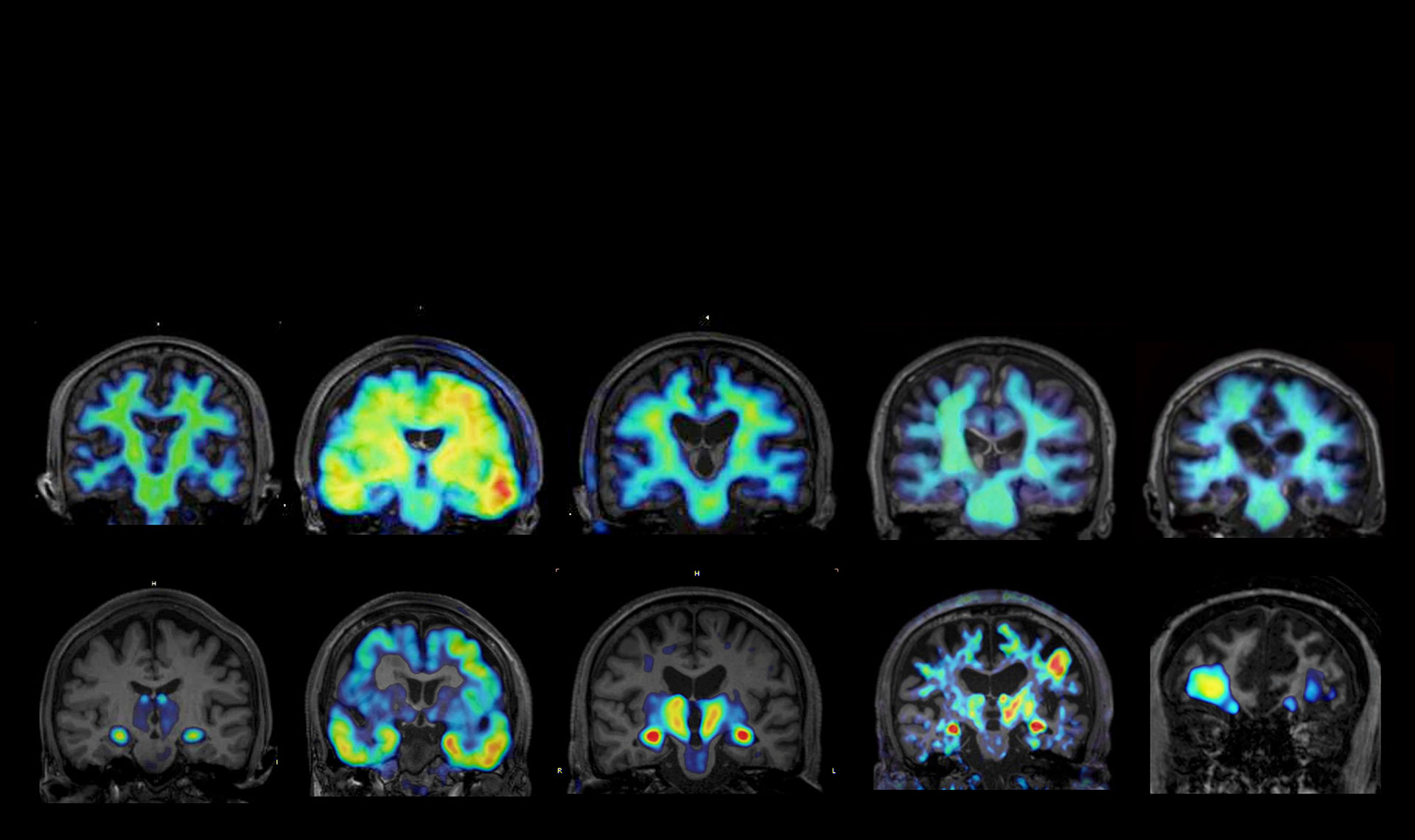
As a research collaborator for Moonshot Goal 7 “Cerebral Seno-inflammation,” I participate under PM Higuchi as an expert in mathematical modeling and AI analysis. In this project, I am responsible for the integrative analysis of multidimensional biological data, including human autopsy brains, model animals, and omics data obtained by other collaborating researchers (Kakita, Seto, Masuda, Hoshino, et al.). Specifically, we aim to identify key substances and origin substances of “seno-inflammation (the linkage between cellular senescence and inflammation)” in dementia pathology using advanced deep learning and causal inference models, and to construct a system for predicting pathological changes through therapeutic intervention. Our ultimate goal is to realize a “digital twin” that proposes optimized prevention and treatment strategies for individual patients by mathematically modeling molecular and cellular mechanisms and integrating them with clinical data.
Research Theme
Development of Mathematical Models for Ultra-Early Prediction and Pathophysiological Elucidation of Dementia Onset
1. Integrative Analysis and Feature Extraction from Multidimensional Biological Data
We are developing analytical methods to integrate multidimensional biological data such as blood tests, imaging examinations, and omics data. In particular, we focus on the integrative analysis of single-cell omics data (single-cell analysis) to extract cell-specific features. Additionally, by developing time-series data analysis methods, we are building the foundation to capture dynamic changes in dementia pathology progression. Through this, we aim to extract disease mechanisms conserved across species as well as human-specific features.
2. Risk Prediction Algorithms Using Deep Learning and Causal Inference
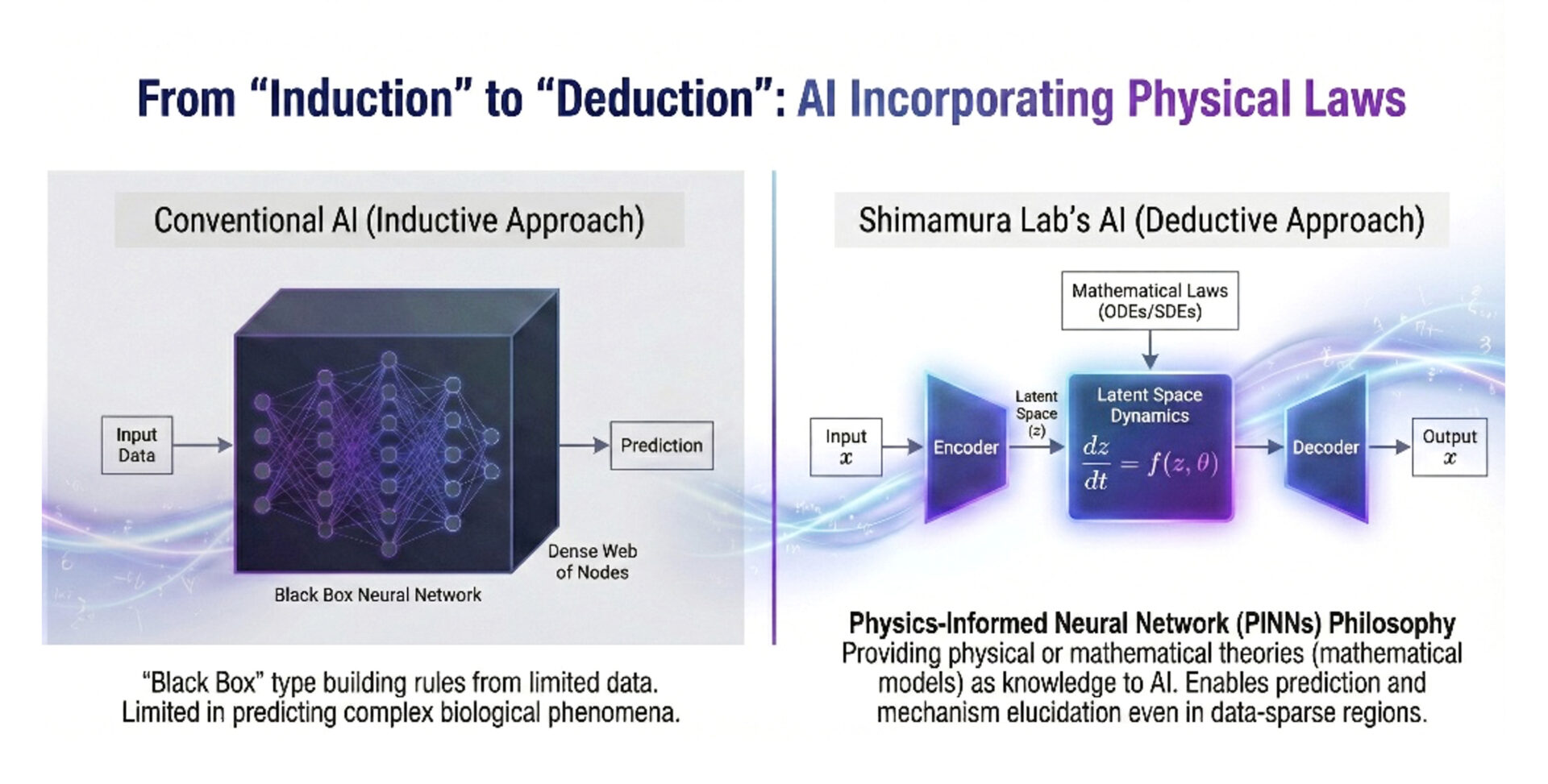
Based on the developed mathematical models, we are working on constructing algorithms to predict dementia onset risk. We develop advanced deep learning models that take clinical data such as blood biomarkers and imaging data as input to predict future cognitive decline risk. Beyond mere prediction, we aim to enhance the interpretability of AI models, which tend to be black boxes, by introducing “causal inference methods” and to visualize the interactions and hierarchical structures among risk factors.
3. Personalized Medicine Simulation Using Digital Twins
In the latter half of the research, we will construct an advanced “digital twin model” that integrates detailed molecular and cellular pathophysiological mechanism models with clinical data. Using this model, it will be possible to simulate the effects of various therapeutic intervention scenarios in virtual space. We will proceed with the development and validation of a “comprehensive decision support system” that performs precise pathophysiological predictions reflecting individual patient biological characteristics and clinical course, and identifies optimal approaches for each patient.
Selected Publications
- Mizukoshi C, Kojima Y, Hayashi S, Abe K, Kasugai D, Shimamura T. scSurv: a deep generative model for single-cell survival analysis. Bioinformatics. 2026 Jan 2; 42(1):btaf671.
- Mizukoshi C, Kojima Y, Nomura S, Hayashi S, Abe K, Shimamura T. DeepKINET: a deep generative model for estimating single-cell RNA splicing and degradation rates. Genome Biol. 2024 Sep 6;25(1):229.
- Majima K, Kojima Y, Minoura K, Abe K, Hirose H, Shimamura T. LineageVAE: reconstructing historical cell states and transcriptomes toward unobserved progenitors. Bioinformatics. 2024 Oct 1;40(10):btae520.
- Kojima Y, Mii S, Hayashi S, Hirose H, Ishikawa M, Akiyama M, Enomoto A, Shimamura T. Single-cell colocalization analysis using a deep generative model. Cell Syst. 2024 Feb 21;15(2):180-192.e7.
- Abe K, Shimamura T. UNMF: a unified nonnegative matrix factorization for multi-dimensional omics data. Brief Bioinform. 2023; 24(5):bbad253.